Complete Genome Sequence of SMBL-WEM22, a Halotolerant Strain of Kosakonia cowanii Isolated from Hong Kong Seawater | Microbiology Resource Announcements
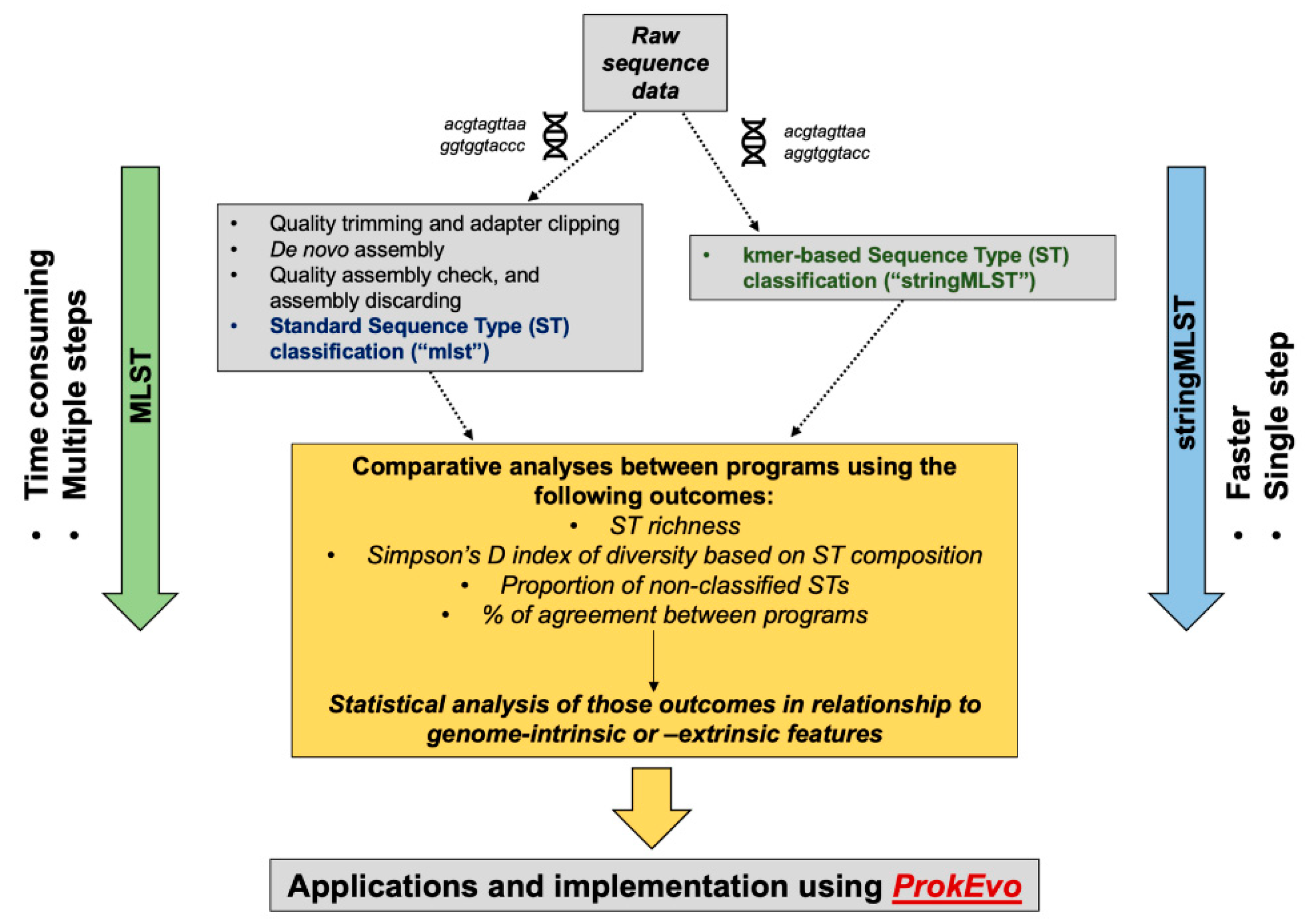
Life | Free Full-Text | Systems-Based Approach for Optimization of Assembly-Free Bacterial MLST Mapping | HTML

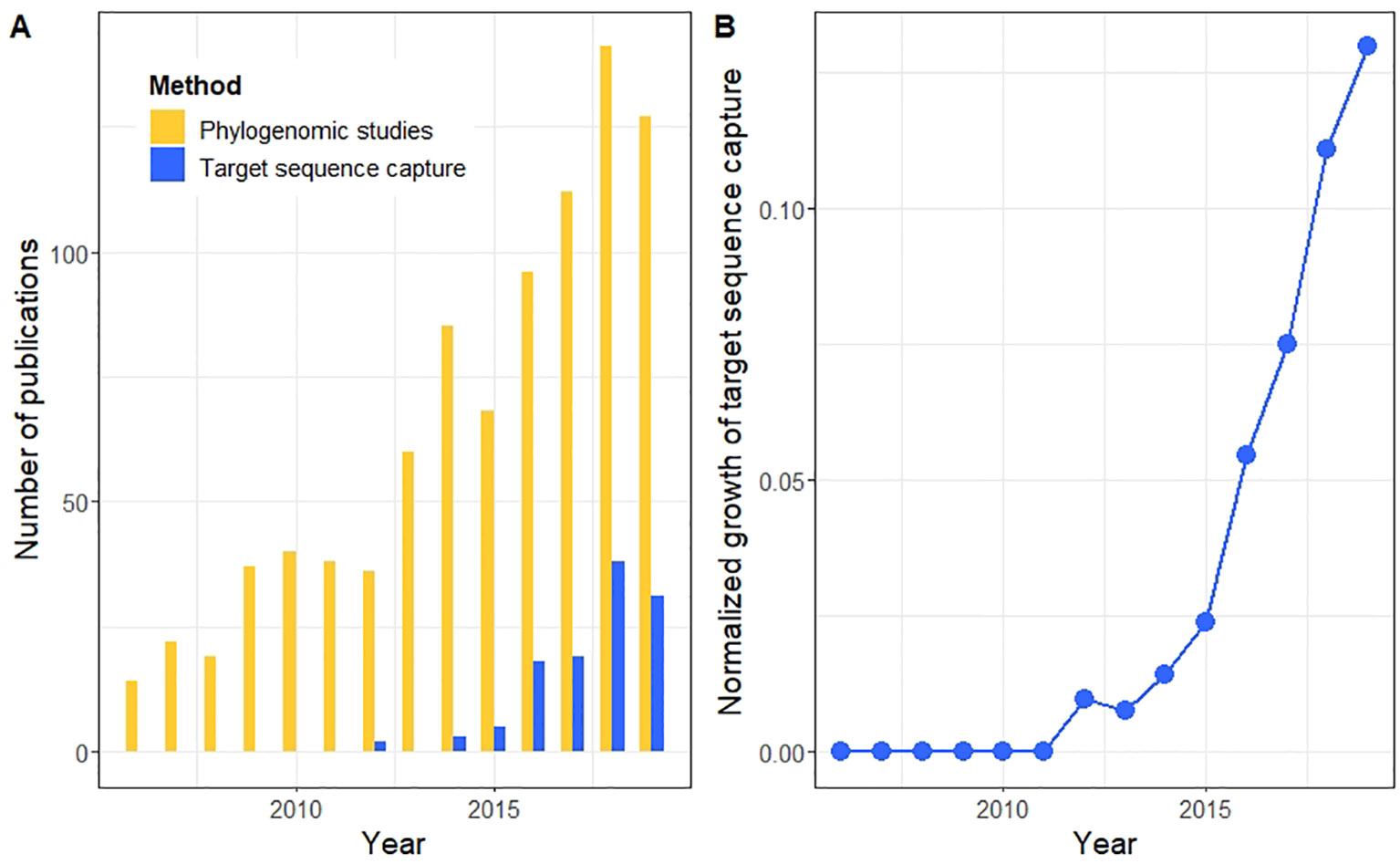
Complementary combination of multiplex high‐throughput DNA sequencing for molecular phylogeny - Suyama - 2022 - Ecological Research - Wiley Online Library
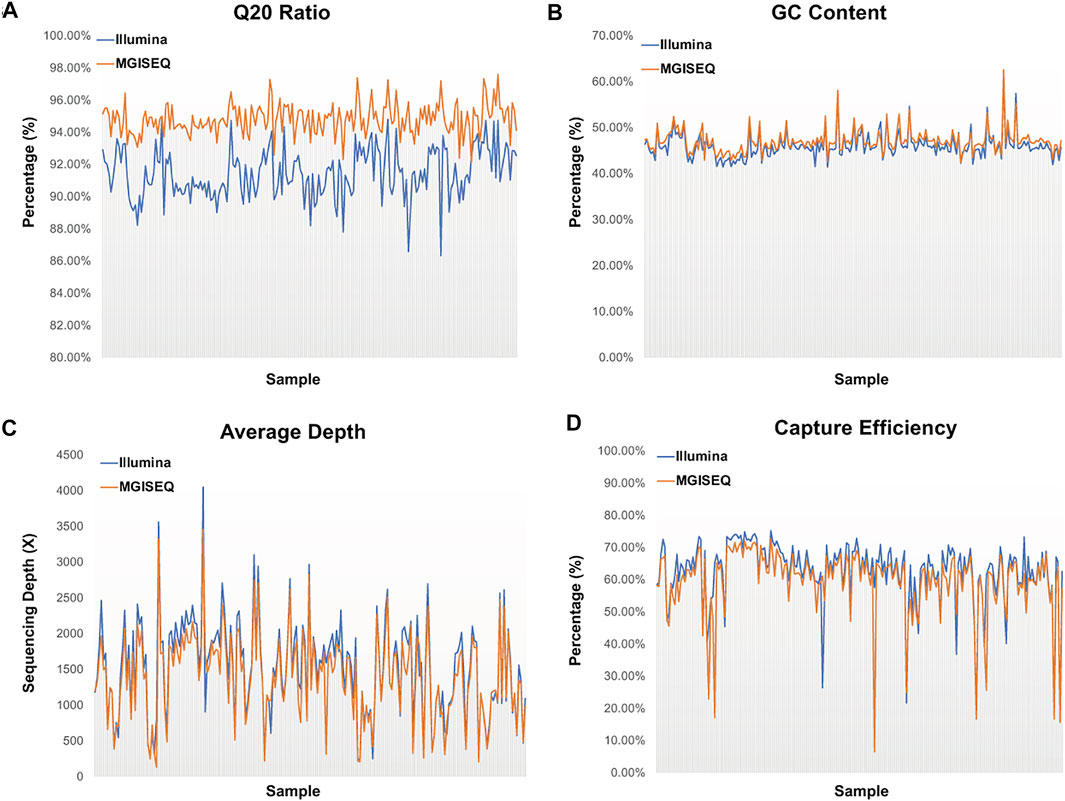
Frontiers | Evaluation of the MGISEQ-2000 Sequencing Platform for Illumina Target Capture Sequencing Libraries | Genetics

Evaluation of WGS performance for bacterial pathogen characterization with the Illumina technology optimized for time-critical situations | Microbiology Society

Assessment of the performance of different imputation methods for low-coverage sequencing in Holstein cattle - Journal of Dairy Science

Microbiome Analysis Laboratory, CNB-CSIC on Twitter: "We are proud to know that FISABIO (Public Health Reasearch Services in Comunidad Valenciana, Spain) has adopted #SqueezeMeta as its reference tool for metagenomic analysis, and #
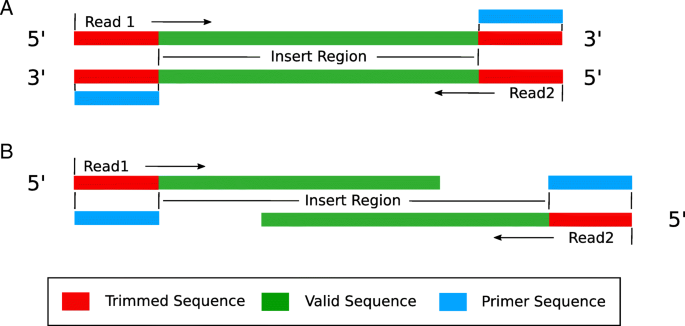
pTrimmer: An efficient tool to trim primers of multiplex deep sequencing data | BMC Bioinformatics | Full Text
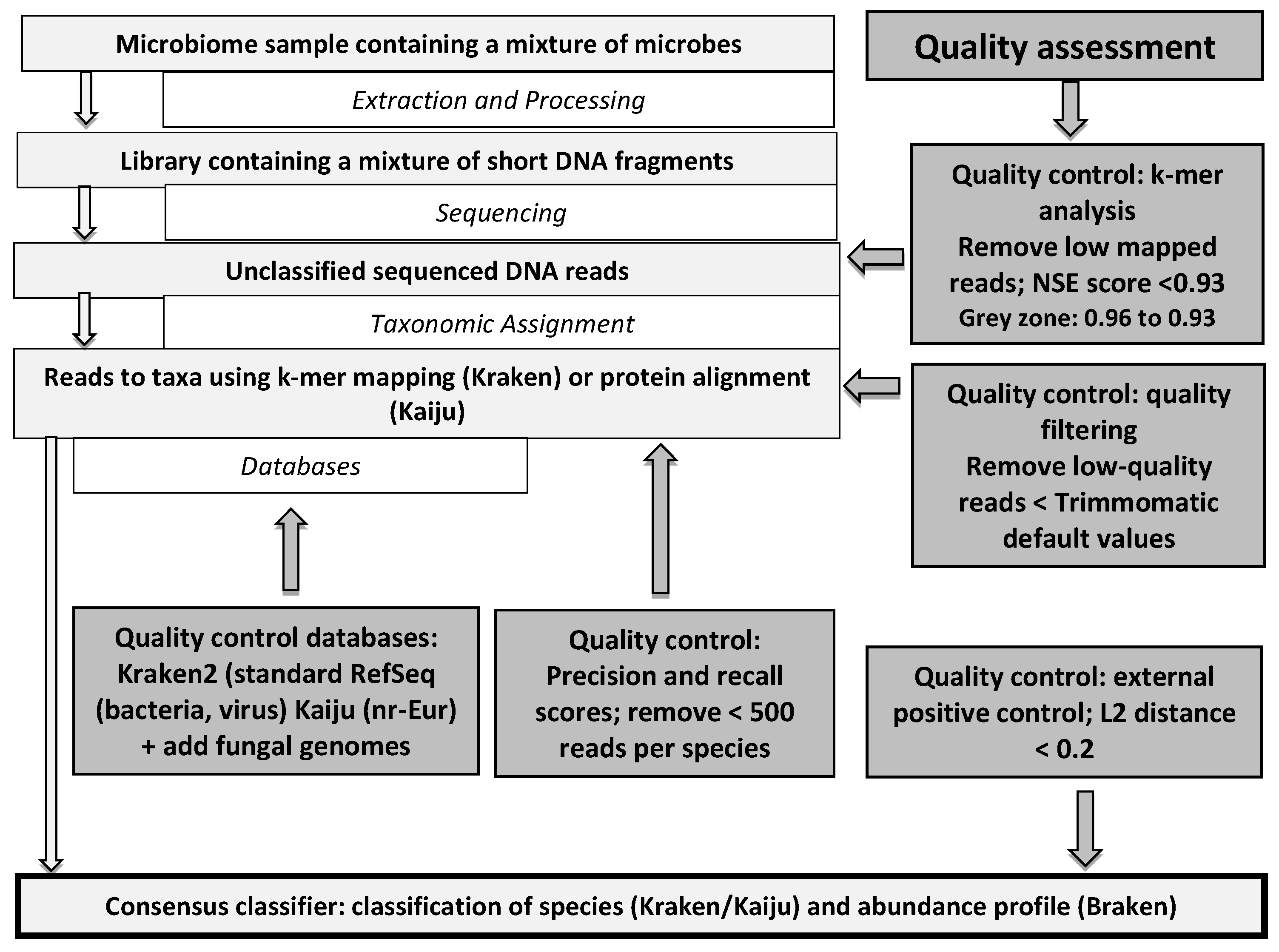

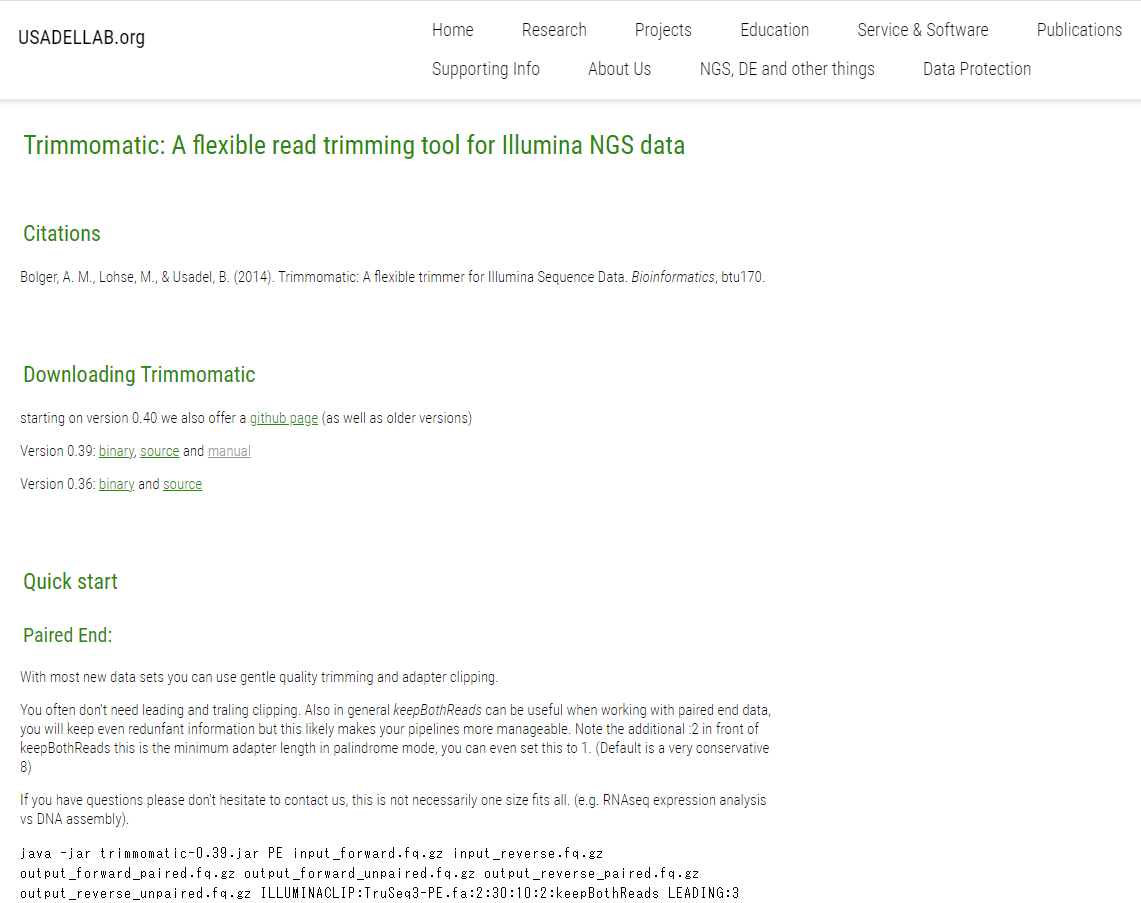




![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/2-Figure2-1.png)

![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/4-Figure3-1.png)
![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/6-Table3-1.png)
![PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar PDF] Trimmomatic: a flexible trimmer for Illumina sequence data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc90910b6e31fe44cddc1e341f21eec0aaa5db44/6-Table2-1.png)